Biology is weird
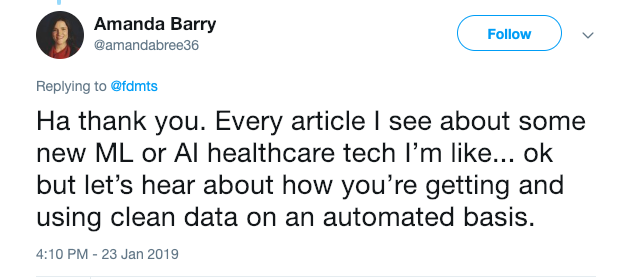
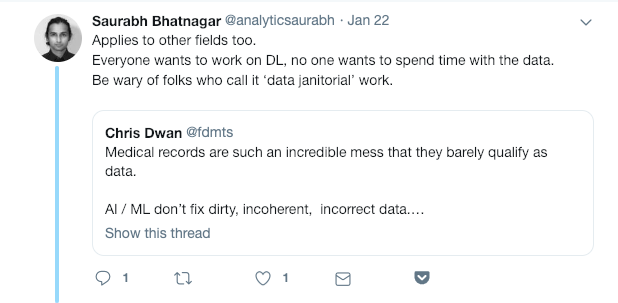
Biology is weird. The data are weird, not least because models evolve rapidly. Today’s textbook headline is tomorrow’s “in some cases,” and next year’s “we used to think.”
It can be hard for non-biologists, particularly tech/math/algorithm/data science/machine learning/AI folks, to really internalize the level of weirdness and uncertainty encoded in biological data.
It is not, contrary to what you have read, anything like the software you’ve worked with in the past. More on that later.
This post is a call for humility among my fellow math / computer science / programmer type people. Relax, roll with it, listen first, come up to speed. Have a coffee with a biologist before yammering about how you’re the first smart person to arrive in their field. You’ll learn something. You’ll also save everybody a bit of time cleaning up your mess.
References
Don’t be the person who walks into a research group meeting carrying a half read copy of “Genome” by Matt Ridley, spouting off about how all you need is to get TensorFlow running on some cloud instances under Lambda and you’re gonna cure cancer.
This is not to speak ill of “Genome,” it’s a great book, and I’m super glad that lots of people have read it – but it no more qualifies you to do the heavy lifting of genomic biology than Lisa Randall’s popular press books prepare you for the mathematical work of quantum physics.
You’ll get more cred with a humble attitude and a well thumbed copy of “Life Ascending” by Nick Lane. For full points, keep Horace Judson’s “The Eighth Day of Creation” on the shelf. Mine rests between Brooks’ “The Mythical Man Month” and “Personality” by Daniel Nettle.
The More Things Change
Back in 2001, the human genome project was wrapping up. One of the big questions of the day was how many genes we would find in the completed genome. First, set aside the important but fundamentally un-answerable question of what, exactly, constitutes a gene. Taking a simplistic and uncontroversial definition, I recall a plurality of well informed people who put the expected total between 100,000 and 200,000.
The answer? Maybe a third to a sixth of that. The private sector effort, published in Science, reported an optimistically specific 26,588 genes. The public effort, published in Nature, reported a satisfyingly broad 30,000 to 40,000.
There was a collective “huh,” followed by the sound of hundreds of computational biologists making strong coffee.
This happens all the time in Biology. We finally get enough data to know that we’ve been holding the old data upside down and backwards.
The fundamental dogma of information flow from DNA to RNA to Protein seems brittle and stodgy when confronted with retroviruses, and honestly a bit quaint in the days of CRISPR. I’ve lost count of the number of lower-case modifiers we have to put on the supposedly inert “messenger molecule” RNA to indicate its various regulatory or even directly bio-active roles in the cell.
Biologists with a few years under their belt are used to taking every observation and dataset with a grain of salt, to constantly going back to basics, and to sighing and making still more coffee when some respected colleague points out that that thing … well … it’s different than we expected.
So no, you’re not going to “cure cancer” by being the first smart person to try applying math to Biology. But you -do- have an opportunity to join a very long line of well meaning smart people who wasted a bunch of time finding subtle patterns in our misunderstandings rather than doing the work of biology, which is to interrogate the underlying systems themselves.
Models
To this day, whenever I look at gene expression pathways I think: “If I saw this crap in a code review, I would send the whole team home for fear of losing my temper.”
My first exposure to bioinformatics was via a seminar series at the University of Michigan in the late 90’s. Up to that point, I had studied mostly computer science and artificial intelligence. I was used to working with human-designed systems. While these systems sometimes exhibited unexpected and downright odd behaviors, it was safe to assume that a plan had, at some point, existed. Some human or group of humans had put the pieces of the system together in a way that made sense to them.
To my eye, gene expression pathways look contrived. It’s all a bit Rube Goldberg down there, with complex and interlocking networks of promotion and inhibition between things with simple names derived from the names of famous professors (and their pets).
My design sensibilities keep wanting to point out that there is no way that this mess is how we work, that this thing needs a solid refactor, and that … dammit … where’s the coffee?
It gets worse when you move from example to example and keep finding that these systems overlap and repeat in the most maddening way. It’s like the very worst sort of spaghetti code, where some crazy global variable serves as the index for a whole bunch of loops in semi-independent pieces of the system, all running in parallel, with an imperfect copy paste as the fundamental unit of editing.
This is what happens when we apply engineering principles to understanding a system that was never engineered in the first place.
Those of us who trained up on human designed systems apply those same subconscious biases that show us a face in the shadows of the moon. We’re frustrated when the underlying model is not based on noses and eyes but rather craters and ridges. We go deep on the latest algorithm or compute system – thinking that surely there’s reason and order and logic if only we dig deep enough.
Biologists roll with it.
They also laugh, stay humble, and drink lots of coffee.